DNA Binding Motif
| Accessions: | ZNF660_2 (HT-SELEX2 May2017), UN0211.1 (JASPAR 2024) |
| Names: | ZNF660 |
| Organisms: | Homo sapiens |
| Libraries: | HT-SELEX2 May2017 1, JASPAR 2024 2 1 Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] 2 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | HT-SELEX |
| Length: | 18 |
| Consensus: | ayTGATkvCCCAyCCTay |
| Weblogo: | 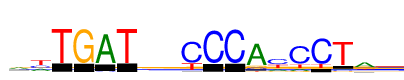 |
| PSSM: | P0 A C G T 01 7679 3303 3528 1193 a 02 4664 4794 742 8773 y 03 0 24 0 15700 T 04 39 0 15692 0 G 05 15300 578 28 146 A 06 3 24 0 15685 T 07 3695 1248 4840 5920 k 08 5761 4151 4132 1659 v 09 183 13769 524 4801 C 10 31 15611 36 131 C 11 73 15697 11 12 C 12 15003 1322 1483 870 A 13 3496 8693 1883 5655 y 14 2073 13506 1062 3197 C 15 28 14847 7 2723 C 16 877 783 774 15179 T 17 9803 1930 4496 1962 a 18 2529 6563 1751 4861 y |
| Type: | Heterodimer |
| Binding TFs: | Q6AZW8 / ZNF660 (Zinc finger, C2H2 type, Zinc-finger double domain, C2H2-type zinc finger, C2H2-type zinc finger) Q6AZW8 |
| Publications: | Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.