DNA Binding Motif
| Accessions: | NR2C2_2 (HT-SELEX2 May2017), MA1536.1 (JASPAR 2024) |
| Names: | NR2C2, NR2C2(var.2) |
| Organisms: | Homo sapiens |
| Libraries: | HT-SELEX2 May2017 1, JASPAR 2024 2 1 Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] 2 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | HT-SELEX |
| Length: | 8 |
| Consensus: | rrGGTCRh |
| Weblogo: | 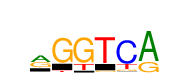 |
| PSSM: | P0 A C G T 01 6236 3111 6420 2861 r 02 10232 719 8395 0 r 03 0 236 18628 1341 G 04 0 0 18628 4790 G 05 0 0 1588 18628 T 06 0 18628 1643 4406 C 07 18628 0 6639 0 R 08 5065 4757 3684 5122 h |
| Type: | Heterodimer |
| Binding TFs: | P49116 (Ligand-binding domain of nuclear hormone receptor, Zinc finger, C4 type (two domains)) NR2C2 (Zinc finger, C4 type (two domains)) P49116 |
| Publications: | Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] Lee H.J, Lee Y.F, Chang C. TR4 orphan receptor represses the human steroid 21-hydroxylase gene expression through the monomeric AGGTCA motif. Biochemical and biophysical research communications 285:1361-8 (2001). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.