DNA Binding Motif
| Accessions: | SNAI3_2 (HT-SELEX2 May2017), MA1559.1 (JASPAR 2024) |
| Names: | SNAI3 |
| Organisms: | Homo sapiens |
| Libraries: | HT-SELEX2 May2017 1, JASPAR 2024 2 1 Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] 2 Rauluseviciute I, Riudavets-Puig R, Blanc-Mathieu R, Castro-Mondragon JA, Ferenc K, Kumar V, Lemma RB, Lucas J, Cheneby J, Baranasic D, Khan A, Fornes O, Gundersen S, Johansen M, Hovig E, Lenhard B, Sandelin A, Wasserman WW, Parcy F, Mathelier A. JASPAR 2024: 20th anniversary of the open-access database of transcription factor binding profiles. Nucleic Acids Res : (2023). [Pubmed] |
| Notes: | HT-SELEX |
| Length: | 10 |
| Consensus: | arCAGGTGya |
| Weblogo: | 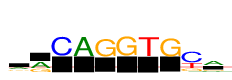 |
| PSSM: | P0 A C G T 01 2952 661 1525 1110 a 02 3917 0 2331 183 r 03 23 6248 0 35 C 04 6248 0 34 3 A 05 118 29 6248 43 G 06 0 18 6248 0 G 07 0 12 28 6248 T 08 94 0 6248 30 G 09 432 6248 1701 3161 y 10 4680 688 1569 553 a |
| Type: | Heterodimer |
| Binding TFs: | Q3KNW1 / SNAI3 (Zinc finger, C2H2 type, Zinc-finger double domain, C2H2-type zinc finger, C2H2-type zinc finger) Q3KNW1 |
| Publications: | Yin Y, Morgunova E, Jolma A, Kaasinen E, Sahu B, Khund-Sayeed S, Das PK, Kivioja T, Dave K, Zhong F, Nitta KR, Taipale M, Popov A, Ginno PA, Domcke S, Yan J, Schübeler D, Vinson C, Taipale J. Impact of cytosine methylation on DNA binding specificities of human transcription factors. Science : (2017). [Pubmed] Bradley CK, Norton CR, Chen Y, Han X, Booth CJ, Yoon JK, Krebs LT, Gridley T. The snail family gene snai3 is not essential for embryogenesis in mice. PLoS One : (2013). [Pubmed] |
Disclaimer and license
These data are available AS IS and at your own risk. The EEAD/CSIC do not give any representation or warranty nor assume any liability or responsibility for the data nor the results posted (whether as to their accuracy, completeness, quality or otherwise). Access to these data is available free of charge for ordinary use in the course of research. Downloaded data have CC-BY-NC-SA license. FootprintDB is also available at RSAT::Plants, part of the INB/ELIXIR-ES resources portfolio.